Màsters i postgraus
Chromatin dynamics in cancer and cell fate
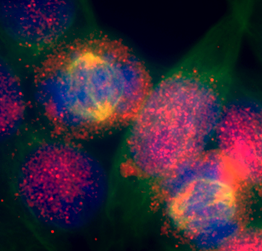
- Our group studies how chromatin factors integrate extracellular signals into the nucleus to regulate gene expression and ultimately, influence cell fate and behavior. Our aim is to understand chromatin dynamics in cancer disease to find novel therapeutic interventions.
- Chromatin alterations are present in different cancer types, thus, their proper identification and understanding of their implication in cancer disease is a first and crucial step towards designing new cures.
We have two projects addressing how chromatin dynamics influence cancer disease.
- Project 1: Chromatin factors as novel biomarkers and drug targets for cancer.
Aims to identify key chromatin factors that shape the cancer epigenetic landscape and how their manipulation can facilitate or impair the access of chemotherapeutic drugs to DNA. We have used an optimized shRNA library (against 900 chromatin factors and a pool approach methodology (figure 1).
Since many epigenetic factors are enzymes that can be targeted by drugs, we expect that using epigenetic drugs in combination with classical chemotherapy could result in synergistic effects that overcome chemoresistance, a fatal outcome for most advanced colorectal cancer patients.
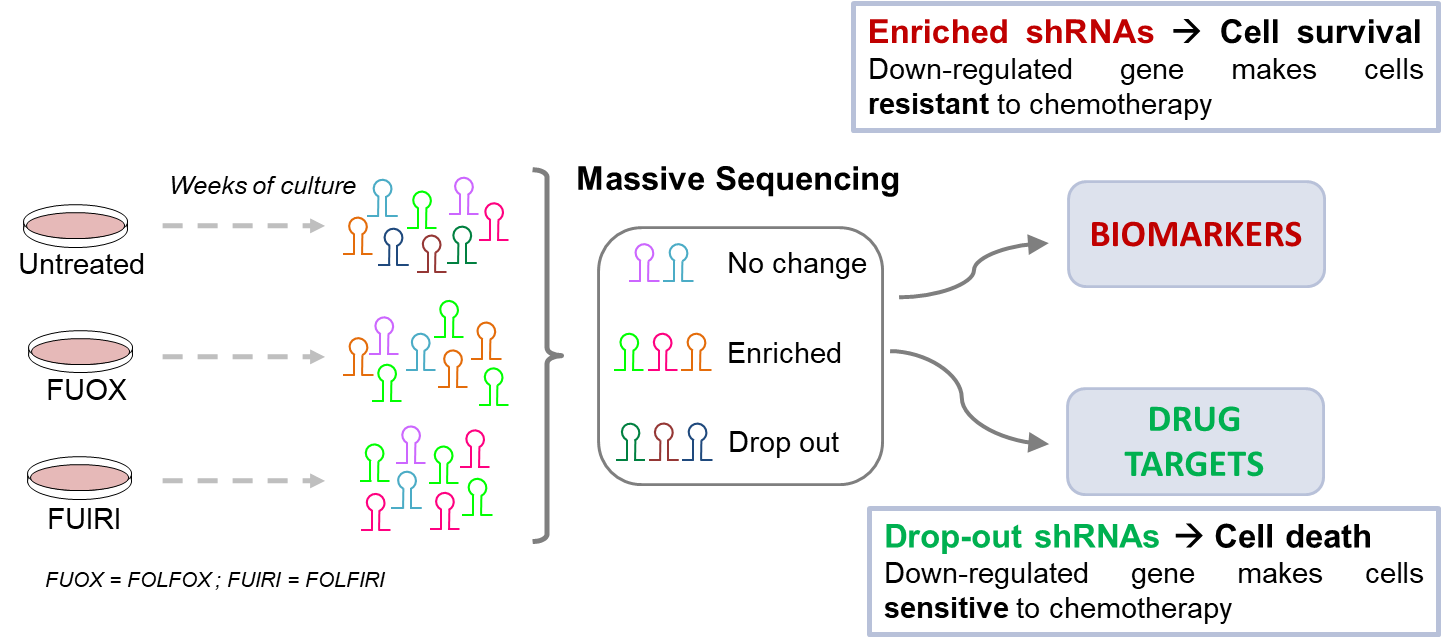
Figure 1. Lost-of-function screen by shRNAs (optimized microRNA backbone described in Cell Rep. 2013 Dec 26;5(6):1704-13) against chromatin factors that resulted in sensitizing to chemotherapy or conferred resistance. Inhibiting these identified factors with epigenetic drugs could synergize with cytotoxic drugs. Modified from PIE16/00011.
- Project 2: Deciphering the junk DNA: a journey through a macrosatellite incRNA.
The other project tackles the effects of DNA demethylation present in repetitive areas of the genome, which occur in most cancer types. This global demethylation associates with genomic instability, but the mechanisms linking both events are not fully understood. We focus our efforts in a particular repetitive sequence, the macrosatellite NBL2, a pericentromeric tandem repeat only present in primates, which is still an enigmatic part of our genome. Upon severe demethylation in a subset of colorectal cancer tumors, NBL2 is de-repressed and transcribed into a lncRNA (TNBL) (figure 2).
The mechanisms by which TNBL contributes to cancer are under intensive investigation in my lab. Moreover, the interplay between active and passive DNA demethylation of structural repetitive areas of the genome historically considered as “Junk DNA” might lead to unprecedented results that bring us closer to successful cancer therapies.
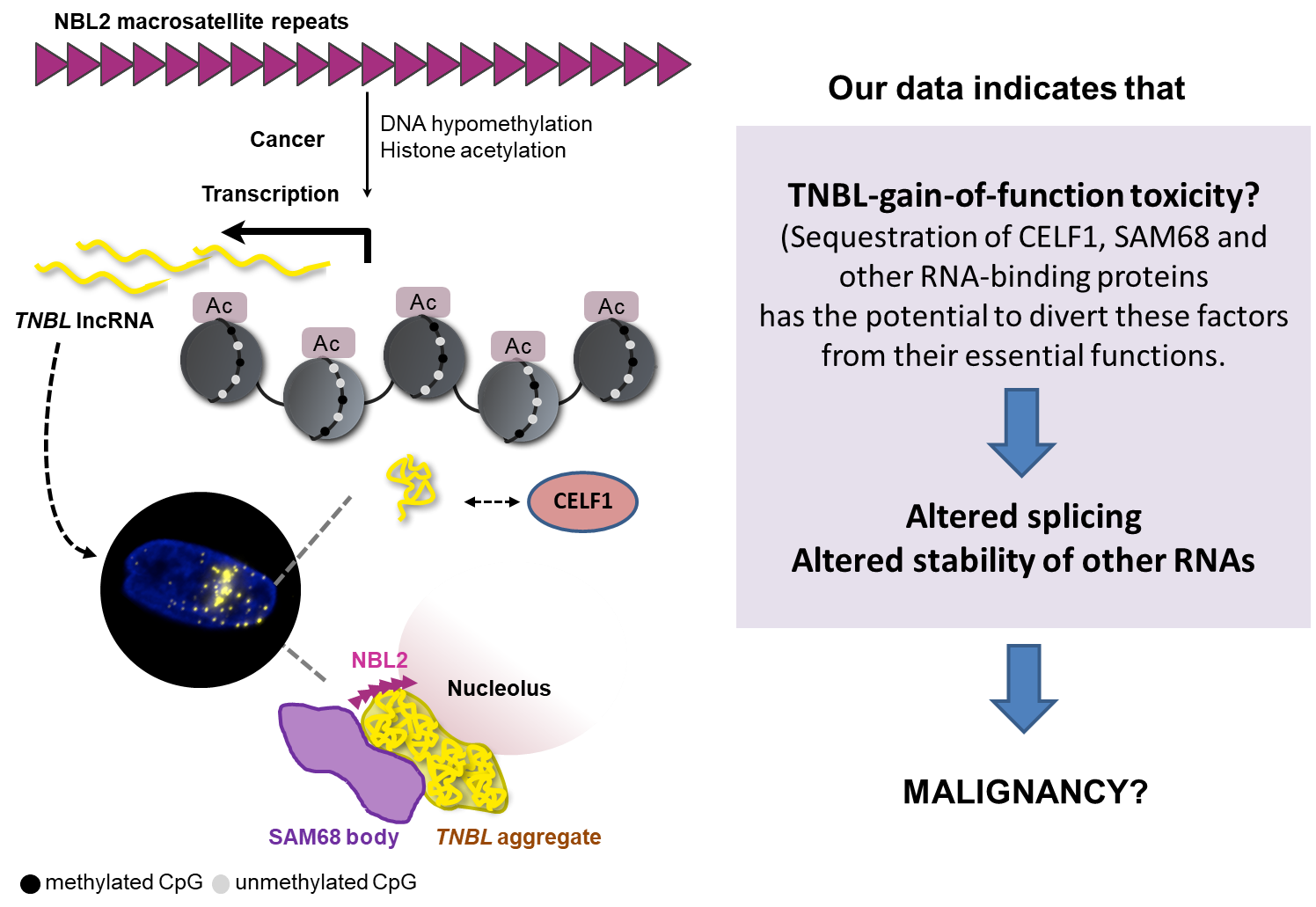
Figure 2. NBL2 epigenetic reprogramming in colorectal cancer leads to unidirectional transcription of a lncRNA that binds to proteins involved in splicing such as CELF1 (dispersed in the nuclear region) and SAM68 (at perinucleolar regions). Modified from Nucleic Acids Res. 2018 Jun 20;46(11):5504-5524.
- Bespalov D, Pino D, Vidal-Guirao S, Franquesa J, Lopez-Ramajo D, Filgaira I, Wan L, O'Sullivan PA, Ley SC, Forcales SV, Rojas JJ, Izquierdo-Serra M, Soler C, Manils J.
Sci Rep. 2024 Oct 3;14(1):22972. doi: 10.1038/s41598-024-74565-4. PMID: 39362963; PMCID: PMC11452207
- Moreta-Moraleda C, Queralt C, Vendrell-Ayats C, Forcales S, Martínez-Balibrea E.
Chromatin factors: Ready to roll as biomarkers in metastatic colorectal cancer? Pharmacol
Res. 2023 Oct;196:106924. doi: 10.1016/j.phrs.2023.106924. Epub 2023 Sep 13. PMID: 37709185
- Zoroddu S, Sanna L, Bordoni V, Lyu W, Murineddu G, Pinna GA, Forcales SV, Sala A, Kelvin DJ, Bagella L.
RNAseq Analysis of Novel 1,3,4-Oxadiazole Chalcogen Analogues Reveals Anti-Tubulin Properties on Cancer Cell Lines.
Int J Mol Sci. 2023 Jul 9;24(14):11263. doi: 10.3390/ijms241411263. PMID: 37511023; PMCID: PMC10379353.
- Solé-Morata N, Baenas I, Etxandi M, Granero R, Forcales SV, Gené M, Barrot C, Gómez-Peña M, Menchón JM, Ramoz N, Gorwood P, Fernández-Aranda F, Jiménez-Murcia S.Solé-Morata N, et al.
The role of neurotrophin genes involved in the vulnerability to gambling disorder.
Sci Rep. 2022 Apr 28;12(1):6925. | doi: 10.1038/s41598-022-10391-w.
- Soler-Botija C, Forcales SV, Bayés Genís A.
Spotlight on epigenetic reprogramming in cardiac regeneration.
Semin Cell Dev Biol. 2019 Apr 26. pii: S1084-9521(18)30240-4.
- Forcales SV, Corominas M.
Chromatin status is key for regeneration.
Semin Cell Dev Biol. 2019. pii: S1084-9521(19)30133-8.
- Sanna L, Piredda R, Marchesi I, Bordoni V, Forcales SV, Calvisi DF, Bagella L.
Chem Biol Interact. 2019;312:108813.
- Dumbovic G, Biayna J, Banús J, Samuelsson J, Roth A, Diederichs S, Alonso S, Buschbeck M, Perucho M, Forcales SV.
Nucleic Acids Res. 2018;46(11):5504-5524.
- Dumbovic G, Forcales SV, Perucho M.
Emerging roles of macrosatellite repeats in genome organization and disease development.
Epigenetics. 2017;12(7):515-526. Review.
- Dumbovic G, Biayna J, Font B, Buschbeck M, Forcales SV.
Barcelona conference on epigenetics and cancer 2016 - beyond cancer genomes.
Epigenetics. 2017;12(3):238-245.
- Cantariño N, Musulén E, Valero V, Peinado MA, Perucho M, Moreno V, Forcales SV, Douet J, Buschbeck M.
Mol Cancer Res. 2016 Sep;14(9):841-8. doi: 10.1158/1541-7786.MCR-16-0034.
- Cantariño N, Fernández-Figueras MT, Valero V, Musulén E, Malinverni R, Granada I, Goldie SJ, Martín-Caballero J, Douet J, Forcales SV, Buschbeck M.
Int J Cancer. 2016;139(5):1106-16.
- Forcales SV.
Potential of adipose-derived stem cells in muscular regenerative therapies.
Front Aging Neurosci. 2015;7:123. eCollection 2015. Review.
- Albini S, Coutinho P, Malecova B, Giordani L, Savchenko A, Forcales SV, Puri PL.
Cell Rep. 2013;3(3):661-70.
- Forcales SV.
The BAF60c-MyoD complex poises chromatin for rapid transcription.
Bioarchitecture. 2012;2(3):104-109.
- Forcales SV, Albini S, Giordani L, Malecova B, Cignolo L, Chernov A, Coutinho P, Saccone V, Consalvi S, Williams R, Wang K, Wu Z, Baranovskaya S, Miller A, Dilworth FJ, Puri PL.
Signal-dependent incorporation of MyoD-BAF60c into Brg1-based SWI/SNF chromatin-remodelling complex.
EMBO J. 20128;31(2):301-16.
- Reseacrh ID (B-2157-2013)
- ORCID (0000-0002-7111-4959)
- Scopus ID: 6507950682

Lejuan Shi (PhD student)
Sonia Forcales (group leader)
Cristina Moreta Moraleda (PhD student)
